Sequence Database Setup: SwissProt
| SwissProt Release 56.0 |
| The syntax of the Fasta title line changed (again!) in SwissProt
release 56.0 (UniProtKB 14.0). This page has been updated to illustrate the new parse rules.
|
|
Overview
SwissProt is a curated protein sequence
database that strives to provide a high level of
annotations (such as the description of the function of a protein, its domains structure,
post-translational modifications, variants, etc.), a minimal level of redundancy and
high level of integration with other databases.
The database is developed by the SWISS-PROT groups at
SIB and
EBI.
Download
Expasy:
ftp://ftp.expasy.org/databases/uniprot/knowledgebase
EBI:
ftp://ftp.ebi.ac.uk/pub/databases/uniprot/knowledgebase
The EBI site mirrors the Expasy site. The files are:
- Version info: reldate.txt
- Fasta file: uniprot_sprot.fasta.gz
- Dat file: uniprot_sprot.dat.gz
To download updates automatically, the relevant definition blocks in
db_update.pl is SwissProt_complete_from_EBI.
There is also a definition for downloading just the SwissProt
Fasta file: SwissProt_fasta_only_from_EBI.
Taxonomy
Taxonomy is predefined in mascot.dat. If you have the Dat file, choose "Swiss-prot DAT".
If you do not wish to have a local copy of the Dat file, and prefer to take taxonomy from the Fasta file instead,
you will need to update the taxonomy definition in mascot.dat. Make a backup copy of mascot.dat,
then use a text editor to change taxonomy block 3 as follows:
# TAXONOMY FOR SwissProt or Trembl from the fasta file
Taxonomy_3
Identifier SwissProt FASTA
Enabled 1 # 0 to disable it
FromRefFile 0
DescriptionLineSep 0 # ctrl a - hex code '1'. For multiple descriptions per entry
SpeciesFiles NCBI:names.dmp
NodesFiles NCBI:nodes.dmp
DefaultRule NCBI, CHOP:W "OS=\(.*\) PE=" # from release 14.0 onwards
end
Note that mascot.dat must be saved as plain text, so be careful if using a word processor,
and ensure the filename is not changed to mascot.dat.txt or something.
Having made these changes, in database maintenance, choose "SwissProt FASTA".
The following taxonomy files are required:
ftp://ftp.ncbi.nih.gov/pub/taxonomy/taxdump.tar.gz
ftp://ftp.ebi.ac.uk/pub/databases/uniprot/knowledgebase/docs/speclist.txt
Note that the taxonomy files go into the taxonomy directory, not into the sequence database
directory. Also, some files need to be unpacked (using tar) as well as uncompressed.
Parse Rules
A typical SwissProt Fasta title line is:
>sp|Q4U9M9|104K_THEAN 104 kDa microneme/rhoptry antigen OS=Theileria annulata GN=TA08425 PE=3 SV=1
You can use either the ID (104K_THEAN) or
the AC (Q4U9M9) as the identifier.
Many people prefer the ID because it is semi-descriptive.
ID from Fasta title: ">..|[^|]*|\([^ ]*\)"
AC from Fasta title: ">..|\([^|]*\)"
Description from Fasta title: ">[^ ]* \(.*\)"
The corresponding line in the Dat file is:
ID 104K_THEAN Reviewed; 893 AA.
ID from Ref file: "^ID \([^ ]*\)"
Configuration
For this first example, the database files were downloaded to
C:\Inetpub\MASCOT\sequence\SwissProt\current,
decompressed using gzip,
and renamed to SwissProt_56.0.dat and SwissProt_56.0.fasta.
When updating an active database, it is important to rename the Fasta file last, because Mascot
will begin database exchange as soon as it sees a new Fasta file that matches the wildcard path for
the database.
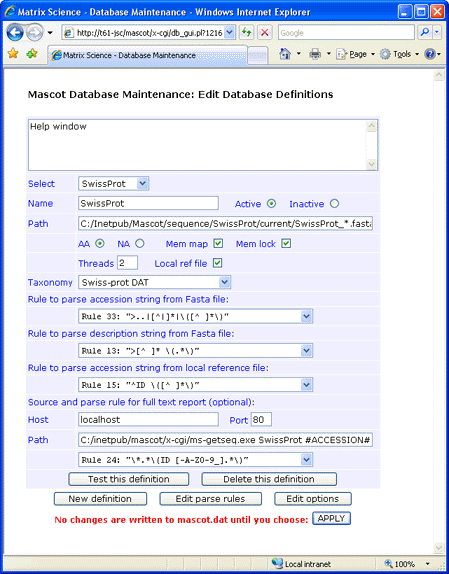
If you decide not to have the reference file locally, full text for individual entries can be retrieved across the web
from an SRS server or Expasy. This can be done using either the ID or the AC as the identifier.
For Expasy, the syntax for the Path field is:
/cgi-bin/get-sprot-raw.pl?#ACCESSION#
Where #ACCESSION# represents either the AC or ID. For an SRS
server, the syntax for the Path field is:
Retrieve by ID: /srsbin/cgi-bin/wgetz?-e+[SWISSPROT-id:#ACCESSION#]+-vn+2
Retrieve by AC: /srsbin/cgi-bin/wgetz?-e+[SWISSPROT-acc:#ACCESSION#]+-vn+2
This screen shot illustrates a configuration in which the identifier is AC and there is no local Dat file:
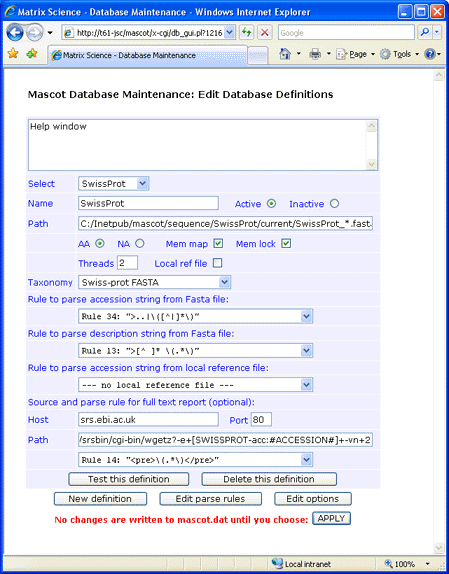
Make sure that the final parse rule has the correct case. Early versions of wgetz return HTML pages tagged
with <PRE>, while later versions use <pre>. Parse rules are always case sensitive.
If you don't require full text in a Mascot Protein View report, simply leave the Host, Port,
and Path fields blank and choose
--- no full text report ---
in the drop down list.
Always test a new definition before applying the changes to mascot.dat.
|