Sequence Database Setup: EMBL EST
Overview
The EST Fasta files from EMBL
contain "single-pass" cDNA sequences, or Expressed Sequence Tags. The sequences are divided
into 10 divisions:
- ENV:Environmental Samples
- FUN:Fungi
- HUM:Human
- INV:Invertebrates
- MAM:Other Mammals
- MUS:Mus musculus
- PLN:Plants
- PRO:Prokaryotes
- ROD:Rodents
- VRT:Other Vertebrates
Download
Individual Fasta files can be downloaded from the
EBI FTP server.
On this help page, the mammals file is used as an example. To work with other divisions, simply substitute the
three letter code. For example, the compressed Fasta file for mammals is em_rel_est_mam.gz,
while the one for fungi is em_rel_est_fun.gz.
Note that versions of wget up to 1.10.x have problems with files larger than 2 GB on 32 bit platforms. The
current stable release, 1.11.4, works correctly.
Windows binaries can be downloaded from Christopher G. Lewis. There is no installer; just unpack the files into a directory
on the system path.
Taxonomy
Taxonomy for EMBL EST files requires Mascot 2.3 or later. For earlier versions of Mascot, configure without
taxonomy. The following taxonomy files are required:
ftp://ftp.ebi.ac.uk/pub/databases/embl/misc/acc_to_taxid.mapping.txt.gz
ftp://ftp.ncbi.nih.gov/pub/taxonomy/taxdump.tar.gz
Note that the taxonomy files go into the taxonomy directory, not into the sequence database
directory. Also, some files need to be unpacked (using tar) as well as uncompressed.
Unigene
The NCBI UniGene
indexes are created by automatically partitioning GenBank sequences into non-redundant sets of
gene-oriented clusters. If UniGene indexes are available locally, results from Mascot searches of
EST databases can be grouped and reported by gene family, rather than by raw EST accession numbers.
To enable UniGene indexes, uncomment the following line, near the top of the
db_update.pl script:
# $local_unigene_directory = "$MASCOT/unigene";
This will cause the required UniGene indexes to be downloaded when the EST databases are next updated.
You will also need to uncomment and possibly modify the relevant lines in the UniGene block of mascot.dat. For example:
arabidopsis /usr/local/mascot/sequence/unigene/arabidopsis/current/At.data
Plants_EST arabidopsis barley maize rice wheat
A control to map database accessions to UniGene families will then be added to the format controls
in MS/MS Summary reports for searches of the enabled databases.
Parse Rules
A typical Fasta title line is:
>EM_EST:AA056886;
AA056886 EST001F Pig Spleen lambda gt 11 Library (Clontech ...
Suitable parse rules are:
Accession from Fasta title: ">EM_EST:\([A-Z0-9]*\);"
Description from Fasta title: ">[^ ]* \(.*\)"
Configuration
For this example, em_rel_est_pln.gz was downloaded to a folder named
C:\sequence\Plants_EST\current.
The file was decompressed using gzip,
and renamed to Plants_EST_103.fasta (because it was from EMBL release 103).
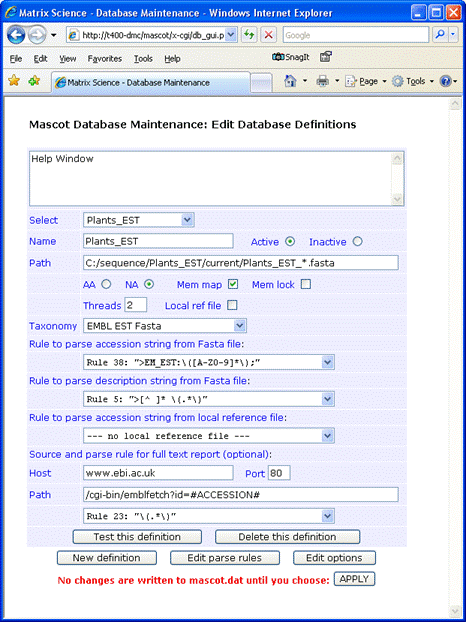
Full text for individual entries can be retrieved across the web
from the EBI at www.ebi.ac.uk. The syntax for the Path field is:
/cgi-bin/emblfetch?id=#ACCESSION#
If you don't require full text in a Mascot Protein View report, simply leave the Host, Port, and Path fields blank
and choose
--- no full text report ---
in the drop down list.
Always test a new definition before applying the changes to mascot.dat.
|